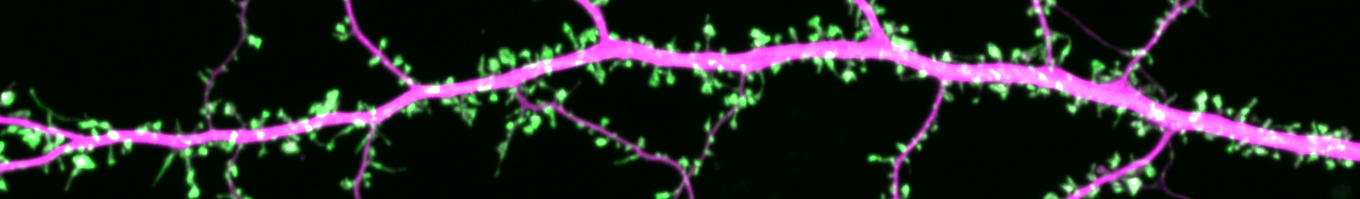
Research in the Blanpied Lab examines protein organization at synapses, to understand mechanisms that control this organization to regulate synaptic transmission.
The dendrites of neurons receive and integrate inputs from hundreds or thousands of partner cells, and most of this input arrives at highly specialized sites called synapses that are distributed over the dendritic tree. Improper regulation of synaptic transmission is implicated in an enormous variety of psychiatric disorders: an imbalance of glutamatergic neurotransmission in particular has been identified in the pathophysiology of diseases ranging from schizophrenia and autism to epilepsy and addiction, and excitatory synapses may be among the earliest targets of Alzheimer’s Disease (AD) pathogenesis.
A key means of synaptic regulation is the control of the number of postsynaptic neurotransmitter receptors available for activation by glutamate release. Thus, understanding the protein trafficking mechanisms involved has broad implications not only for understanding the etiology of many diseases but more generally for defining the cellular basis of nervous system function and disorder.
We focus on processes that operate rapidly and very locally to regulate synaptic function on a moment-to-moment basis. These mechanisms are of particular interest because they likely underlie the initial stages of memory formation, and their perturbation produces severe and long-lasting damage to synaptic and neuronal morphology. Mechanisms we are studying currently include the organization and dynamics of the proteins at each synapse that control receptor number and position at the synapse, the continuous internalization and recycling of synaptic receptors, the postsynaptic actin cytoskeleton that controls postsynaptic morphology and regulates receptor trafficking in many ways, and the trans-synaptic molecular bridges that align presynaptic nerve terminals with postsynaptic specializations. A key finding has been that important aspects of presynaptic and postsynaptic function are well aligned with each other across the synaptic cleft, in a structure we termed the “nanocolumn.”

Understanding the complex set of molecular reactions that underlie endocytosis and other trafficking events requires examining these events in live cells. Thus, the lab has turned to high-resolution, high-speed, multi-color confocal and super-resolution imaging of proteins in cultured neurons. These approaches provide a way of examining the kinetics and spatial features of the molecular events at individual living synapses. We utilize a number of novel assays centered on single-molecule localization and tracking using Photoactivated Localization Microscopy (PALM) to examine protein positioning and movement at the synapse in living cells at the highest resolution possible. These tools provide the perfect complement to whole-cell and single-channel electrophysiological assays, which naturally track receptor function at high speed but typically with little or no spatial resolution. The lab aims to combine these approaches with molecular perturbation of protein function to provide unprecedented spatial, temporal, and molecular understanding of the protein trafficking events in spines.
Support
We are grateful to a number of sources for supporting our research. Our principle support is from the NIMH through a MERIT award R37 MH080046. We also collaborate with Steve Meriney at Pitt and CMU (CRCNS/NINDS), James Trimmer at UC Davis, with Saima Riazuddin and Zubair Ahmed here in the Department of Otorhinolaryngology, and with Thomas Biederer at Yale University (R01 MH119826) and are grateful for past support through the CRCNS mechanism in a collaborative R01 that we shared with Sri Raghavachari at Duke University/UMD College Park, and the BRAIN Initiative with Megan Rizzo and Andrea Meredith. The Dana Foundation, the Broad Foundation, and the Carski Foundation provided important support for the development of PALM in the lab. We also are deeply thankful that the Kahlert Foundation has been able to support the Department and our lab.